FANCI as a candidate ovarian cancer-predisposing gene
Fierheller CT, Guitton-Sert L, Alenezi WM, Revil T, Oros KK, et al. A functionally impaired missense variant identified in French Canadian families implicates FANCI as a candidate ovarian cancer-predisposing gene. Genome Medicine (2021)
How they used Xena
They downloaded data from Xena to run a KM survival analysis stratifying ovarian cancer patients by FANCI expression.
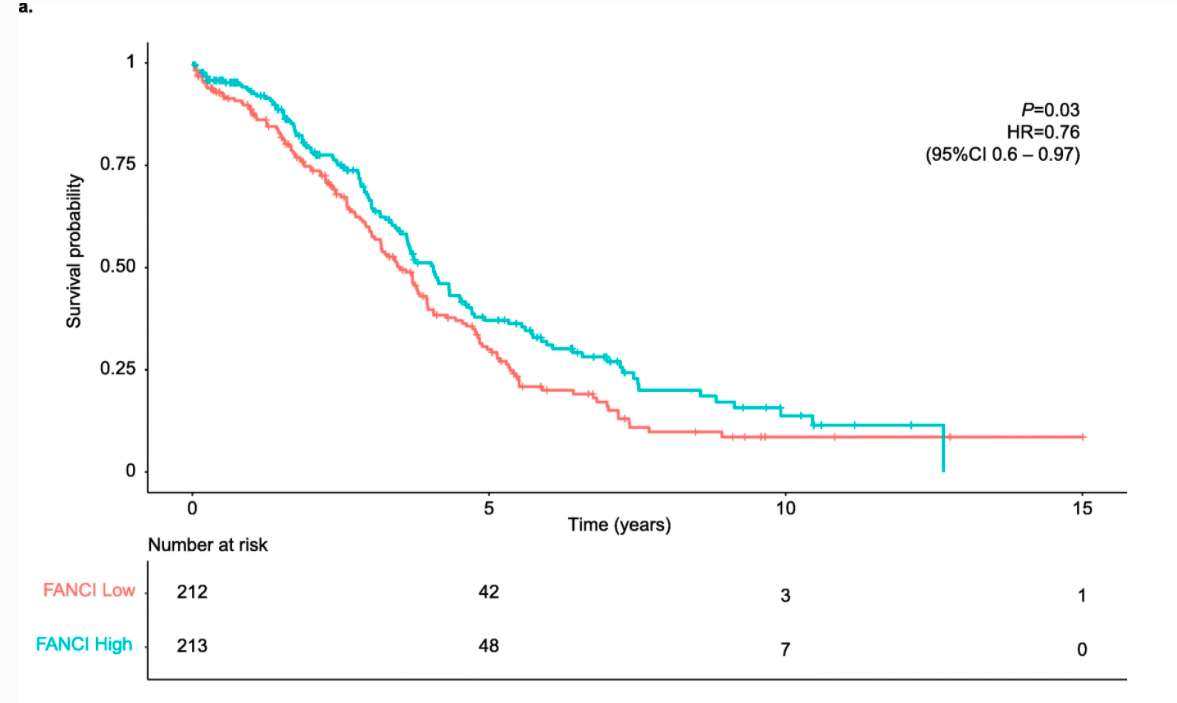
Portion of Figure 5. KM plot made using data from Xena.
Join the conversation on Twitter about this publication
This paper has been highlighted on the podcast Health Matters as well as on BioNews!
Paper
Background
Familial ovarian cancer (OC) cases not harbouring pathogenic variants in either of the BRCA1 and BRCA2 OC-predisposing genes, which function in homologous recombination (HR) of DNA, could involve pathogenic variants in other DNA repair pathway genes.
Methods
Whole exome sequencing was used to identify rare variants in HR genes in a BRCA1 and BRCA2 pathogenic variant negative OC family of French Canadian (FC) ancestry, a population exhibiting genetic drift. OC cases and cancer-free individuals from FC and non-FC populations were investigated for carrier frequency of FANCI c.1813C>T; p.L605F, the top-ranking candidate. Gene and protein expression were investigated in cancer cell lines and tissue microarrays, respectively.
Results
In FC subjects, c.1813C>T was more common in familial (7.1%, 3/42) than sporadic (1.6%, 7/439) OC cases (P = 0.048). Carriers were detected in 2.5% (74/2950) of cancer-free females though female/male carriers were more likely to have a first-degree relative with OC (121/5249, 2.3%; Spearman correlation = 0.037; P = 0.011), suggesting a role in risk. Many of the cancer-free females had host factors known to reduce risk to OC which could influence cancer risk in this population. There was an increased carrier frequency of FANCI c.1813C>T in BRCA1 and BRCA2 pathogenic variant negative OC families, when including the discovery family, compared to cancer-free females (3/23, 13%; OR = 5.8; 95%CI = 1.7–19; P = 0.005). In non-FC subjects, 10 candidate FANCI variants were identified in 4.1% (21/516) of Australian OC cases negative for pathogenic variants in BRCA1 and BRCA2, including 10 carriers of FANCI c.1813C>T. Candidate variants were significantly more common in familial OC than in sporadic OC (P = 0.04). Localization of FANCD2, part of the FANCI-FANCD2 (ID2) binding complex in the Fanconi anaemia (FA) pathway, to sites of induced DNA damage was severely impeded in cells expressing the p.L605F isoform. This isoform was expressed at a reduced level, destabilized by DNA damaging agent treatment in both HeLa and OC cell lines, and exhibited sensitivity to cisplatin but not to a poly (ADP-ribose) polymerase inhibitor. By tissue microarray analyses, FANCI protein was consistently expressed in fallopian tube epithelial cells and only expressed at low-to-moderate levels in 88% (83/94) of OC samples.
Conclusions
This is the first study to describe candidate OC variants in FANCI, a member of the ID2 complex of the FA DNA repair pathway. Our data suggest that pathogenic FANCI variants may modify OC risk in cancer families.